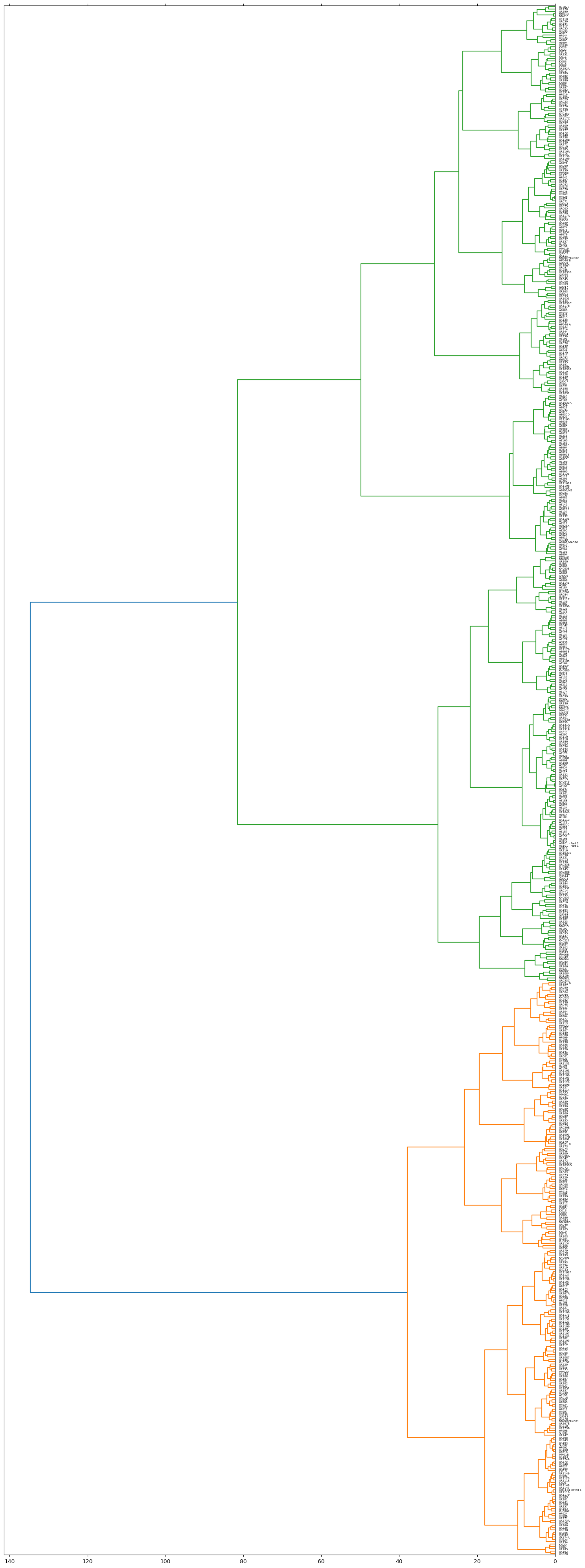
0 : ['AS011', 'AS018', 'AS020', 'AS029', 'AS035C', 'AS036', 'AS037', 'AS041', 'AS050', 'AS054', 'AS063B', 'AS065', 'AS066', 'AS083', 'AS101 - Part 1', 'AS101 - Part 2', 'AS110', 'AS111', 'AS115', 'AS122', 'AS125', 'AS133', 'AS134', 'AS137', 'AS164', 'AS168', 'AS177', 'AS178', 'AS183', 'AS188', 'AS192', 'AS197', 'AS200', 'AS206', 'AS35B', 'HP052', 'HP056', 'UR013', 'UR053B', 'UR053D', 'UR053E', 'UR058A', 'UR058B', 'UR094', 'UR095', 'UR098', 'UR099', 'UR100', 'UR101', 'UR1033E', 'UR1040', 'UR1105', 'UR1113', 'UR1116', 'UR1141', 'UR1150', 'UR1176', 'UR121', 'UR125', 'UR131B', 'UR131C', 'UR136', 'UR137', 'UR142', 'UR145', 'UR161', 'UR186', 'UR187', 'UR194', 'UR216', 'UR223', 'UR247', 'AS001', 'AS002', 'AS003', 'AS006', 'AS007', 'AS008', 'AS009', 'AS014', 'AS063', 'AS072', 'AS073', 'AS156', 'AS170', 'AS187', 'AS209', 'CM009', 'HP037', 'HP039', 'HP047', 'HP053', 'KH0049', 'KH0057', 'KH0058', 'KH0081', 'KH0083', 'MM002', 'MM004', 'MM008', 'MM009', 'MM010', 'MM012', 'MM014', 'MM015', 'MM016', 'MM017', 'QU003', 'QU005', 'QU011', 'QU012', 'QU014', 'QU019', 'UR002', 'UR011', 'UR014', 'UR015', 'UR049', 'UR053A', 'UR085', 'UR104', 'UR108', 'UR118', 'UR119', 'UR122', 'UR130', 'UR131A', 'UR143', 'UR166', 'UR253'] ,
1 : ['AS194', 'AS195', 'AS196', 'AS199', 'HP003', 'HP005', 'HP012', 'HP013', 'HP018', 'HP021', 'HP022', 'HP023', 'UR005', 'UR008', 'UR010', 'UR033', 'UR034', 'UR061', 'UR062', 'UR086', 'UR090', 'UR1033G', 'UR105', 'UR1051', 'UR107', 'UR1084', 'UR1087', 'UR1100', 'UR1102', 'UR1103', 'UR1107', 'UR1109', 'UR1114', 'UR1124', 'UR1126', 'UR1127', 'UR1135', 'UR1136', 'UR1140', 'UR1144', 'UR1147', 'UR1154', 'UR1161', 'UR1162B', 'UR1165', 'UR1167', 'UR1175', 'UR120', 'UR127', 'UR150', 'UR151', 'UR152', 'UR174', 'UR192', 'UR197', 'UR202', 'UR204', 'UR207', 'UR208', 'UR212', 'UR213', 'UR217', 'UR219', 'UR220', 'UR222', 'UR225', 'UR227', 'UR228', 'UR229', 'UR232', 'UR233', 'UR240', 'UR246', 'UR294', 'HP009', 'HP017', 'HP034', 'HP055', 'JC001', 'JC017', 'KH0001', 'KH0032', 'MM006/AN001', 'MM020', 'MM022', 'MM1086', 'QU016', 'UR004', 'UR017', 'UR018', 'UR022', 'UR088', 'UR113', 'UR1131', 'UR1138', 'UR138', 'UR141', 'UR146', 'UR149', 'UR154', 'UR193', 'UR206', 'UR221', 'UR237', 'UR255', 'UR258', 'UR260', 'UR261', 'UR262', 'UR264', 'UR267A', 'UR270', 'UR277', 'UR279', 'UR280', 'UR293'] ,
2 : ['AS010', 'AS013', 'AS015', 'AS021', 'AS023', 'AS026A', 'AS026B', 'AS035D', 'AS039', 'AS043', 'AS048', 'AS059', 'AS060', 'AS064', 'AS065B', 'AS077', 'AS085', 'AS089', 'AS092', 'AS093', 'AS112', 'AS128', 'AS132', 'AS142', 'AS153', 'AS155', 'AS157', 'AS158', 'AS159', 'AS160', 'AS169', 'AS171', 'AS173', 'AS174', 'AS182', 'AS185', 'AS186', 'AS189', 'AS201', 'AS202', 'AS203', 'AS205', 'AS207A', 'AS207C', 'AS210', 'AS211', 'AS212', 'AS213', 'AS214', 'AS215', 'AS35A', 'UR040', 'UR041', 'UR042', 'UR051', 'UR1033A', 'UR1097', 'UR1108', 'UR1121', 'UR1146', 'UR1151', 'UR1162A', 'UR1163', 'UR1166', 'UR132', 'AS012', 'AS016', 'AS019', 'AS024', 'AS027', 'AS028', 'AS044', 'AS045', 'AS069', 'AS071', 'AS090/N2', 'AS094', 'AS207B', 'KH0080', 'UR1034', 'UR1098'] ,
3 : ['HP001', 'HP002', 'HP006', 'HP007', 'HP008', 'HP010', 'HP011', 'HP024', 'HP029', 'HP030', 'UR020', 'UR038', 'UR046', 'UR1104', 'UR1106', 'UR1118', 'UR1119', 'UR1120', 'UR1123', 'UR1124 Detail 1', 'UR1143', 'UR1145', 'UR1148', 'UR1152', 'UR1179', 'UR1180', 'UR129', 'UR162', 'UR183', 'UR185', 'UR200', 'UR218', 'UR226', 'UR234', 'UR248', 'UR249', 'UR274B', 'HP027', 'JC007', 'JC018', 'JC020', 'JC021', 'KH0067', 'KH0197', 'MM018', 'QU010', 'QU015', 'SH001', 'SH002', 'UR001', 'UR026', 'UR1149', 'UR147', 'UR153', 'UR164', 'UR165', 'UR188', 'UR201', 'UR256', 'UR257', 'UR266', 'UR267B', 'UR268', 'UR269', 'UR271', 'UR273A', 'UR273B', 'UR274A', 'UR275', 'UR278'] ,
4 : ['UR103', 'UR180', 'UR282', 'UR283', 'UR285', 'UR286', 'UR287', 'UR289', 'UR290', 'JC002', 'JC003', 'JC004', 'JC005', 'JC006', 'JC008', 'JC009', 'JC010', 'JC011', 'JC012', 'JC013', 'JC014', 'JC015', 'JC016', 'JC019', 'JC022', 'JC023', 'UR251', 'UR284', 'UR288', 'UR291A', 'UR292A'] ,
5 : ['AS017', 'AS061/MA036', 'AS062', 'AS204', 'AS215F', 'HP004', 'UR007', 'UR016', 'UR019', 'UR023', 'UR024', 'UR029', 'UR045', 'UR057', 'UR077', 'UR096', 'UR097', 'UR1052', 'UR109', 'UR1130', 'UR116A', 'UR116B', 'UR117C', 'UR148', 'UR156', 'UR175', 'UR205', 'UR215', 'UR230', 'UR238', 'UR241', 'AS005', 'HP016', 'HP038', 'HP044', 'KH0033', 'KH0350', 'UR003', 'UR012', 'UR169', 'UR177', 'UR184', 'UR196', 'UR254', 'UR272', 'UR276'] ,
6 : ['AS076', 'AS191', 'HP015', 'HP020', 'HP032', 'HP051 B', 'UR006', 'UR021', 'UR027', 'UR053C', 'UR056B', 'UR066', 'UR067', 'UR069', 'UR078', 'UR079', 'UR082', 'UR091', 'UR093', 'UR102', 'UR1032', 'UR1033C', 'UR1033F', 'UR1053', 'UR1058', 'UR106', 'UR1091', 'UR1096', 'UR114', 'UR117B', 'UR117D', 'UR124', 'UR128', 'UR134', 'UR135', 'UR139', 'UR140', 'UR160', 'UR170', 'UR173', 'UR179', 'UR189', 'UR195', 'UR203', 'UR210', 'UR214', 'UR235', 'UR239', 'UR242', 'UR243', 'UR244', 'UR250', 'AS080', 'HP033', 'HP040', 'HP045', 'HP046 A', 'HP048', 'HP051 A', 'HP057', 'KH0227', 'MM001', 'MM003', 'MM021', 'QU004', 'QU007', 'QU009', 'QU013', 'QU017', 'QU021', 'UR059', 'UR089', 'UR1095', 'UR155', 'UR190', 'UR191', 'UR198', 'UR211', 'UR231', 'UR252'] ,
7 : ['AS025', 'AS055', 'AS081', 'AS082', 'AS129', 'AS139', 'AS172', 'AS184', 'HP050', 'UR084', 'UR1099', 'UR1117', 'UR178', 'UR182', 'AS004', 'AS042', 'AS182B', 'KH0267', 'MM011', 'MM013', 'UR039', 'UR050', 'UR052', 'UR054', 'UR055', 'UR110', 'UR112', 'UR144'] ,
8 : ['AS075', 'AS078', 'AS079', 'AS193', 'AS198', 'HP014', 'HP019', 'HP025', 'HP026', 'HP028', 'HP031', 'HP049', 'UR009', 'UR028', 'UR037', 'UR048', 'UR056A', 'UR056C', 'UR060', 'UR064', 'UR065', 'UR068', 'UR070', 'UR071', 'UR072', 'UR073', 'UR075', 'UR076', 'UR080', 'UR081', 'UR087', 'UR092', 'UR1033B', 'UR1033D', 'UR1049', 'UR1088', 'UR111', 'UR117A', 'UR123', 'UR157', 'UR158', 'UR159', 'UR168', 'UR171', 'UR172', 'UR199', 'UR245', 'UR265', 'AS074', 'HP042', 'HP043', 'HP046 B', 'HP054', 'MM005', 'MM007/AN002', 'MM019', 'QU001', 'QU002', 'QU006', 'QU008', 'QU018', 'QU020', 'QU022', 'UR032', 'UR047', 'UR063', 'UR074', 'UR1057', 'UR167', 'UR209', 'UR259', 'UR263'] ,